
PIR Resources for Protein Knowledge Discovery Through BioMart web services as an on-going initiative by the EMBRACE project. The unified presentation and interoperability of these diverse resources is made possible Large quantities of biological data generated byĭifferent research centres have been exposed with a variety of programmatic interfaces. Fly, chicken and Arabidopsis Reactome databases are under development. Tools for high-throughput data analysis and data mining, and data are provided. 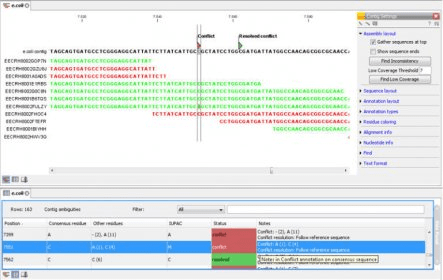
Orthology-based inference of reactions is available for 22 species. Pathways are authored and peer-reviewed by domain experts. Reactome is a curated, open source database of human biological processes. Presented by: Gopal Gopinathrao, Cold Spring Harbor Laboratory, US InterPro integrates data from ten signature databases and five structural databases, providing information on protein families, domains, repeats and sites in which identifiable features found in known proteins can be applied to novel protein sequences. This demonstration explores the wealth of protein annotation data available through the INTERPRO database ( ). Presented by: Jennifer McDowall, EMBL-EBI, UK Join us for a tour through the multitude of toolsĪvailable in PredictProtein as we show you how to leverage the service in your day-to-day work. PredictProtein is a comprehensive sequenceĪnalysis server that incorporates an increasing number of services and databases Presented by: Guy Yachdav, Laszlo Kajan, Columbia University, US PredictProtein - Beyond Secondary Structure Prediction To design complex genetic constructs that will be presented through a software demonstration. The GenoCAD environment (provides a structured and rigorous methodology for organizations We are creating the infrastructure making it possible for non-specialist to design large genetic constructs used in basicīiological research or product development programs. Presented by: Jean Peccoud, Virginia Tech, US These tools include gene expression analysis, primer design, molecular cloning, phylogenetic analyses, and more. For a comprehensive list of features and capabilities, visit the vendor website.GenoCAD: Computer Assisted Design of genetic constructs Built upon the Genomics Workbench framework, this software has been optimized for use with samples from humans or a number of model organisms. To learn more, visit the vendor website.ĬLC Main Workbench contains a variety of toolkits to work with DNA, RNA, and protein analysis. Its main features include the ability to quickly analyze complex data, modify or personalize workflows, filter and visualize your data, and compare results with relevant databases. CLC Genomics Workbench supports key next-generation sequencing features within genomics, transcriptomics, and epigenomics, and includes all the tools of CLC Main Workbench. To learn more, visit the vendor website.ĬLC Biomedical Workbench offers flexible, ready-to-use analysis workflows. CLC Genomics Workbench incorporates cutting-edge technology and algorithms, while also supporting and integrating with the rest of a typical NGS workflow. Its user-friendly and intuitive interface essentially takes high-throughput analysis solely from bioinformatics programmers doing command-line scripts, and opens it up to scientists searching for biological results. 
CLC Genomics Workbench enables researchers to rapidly analyze and visualize large amounts of data generated by NGS machines. CLC Genomics Workbench is designed to solve the data analysis challenges of high-throughput sequencing with high-throughput sequencing machines.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
